import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt5 first 1000
input_families = "../exp/20240506_test_with_first1000/results/TAIR10.first1000.mcl.tsv"
families_df = pd.read_csv(input_families, sep="\t", names=["gene", "family"])
families_df.head()| gene | family | |
|---|---|---|
| 0 | AT1G14930 | 1 |
| 1 | AT1G14940 | 1 |
| 2 | AT1G14950 | 1 |
| 3 | AT1G14960 | 1 |
| 4 | AT1G23120 | 1 |
families_count = families_df.groupby("family").size()
families_countfamily
1 6
2 6
3 5
4 5
5 5
..
96 2
97 2
98 2
99 2
100 1
Length: 100, dtype: int64sns.histplot(families_count)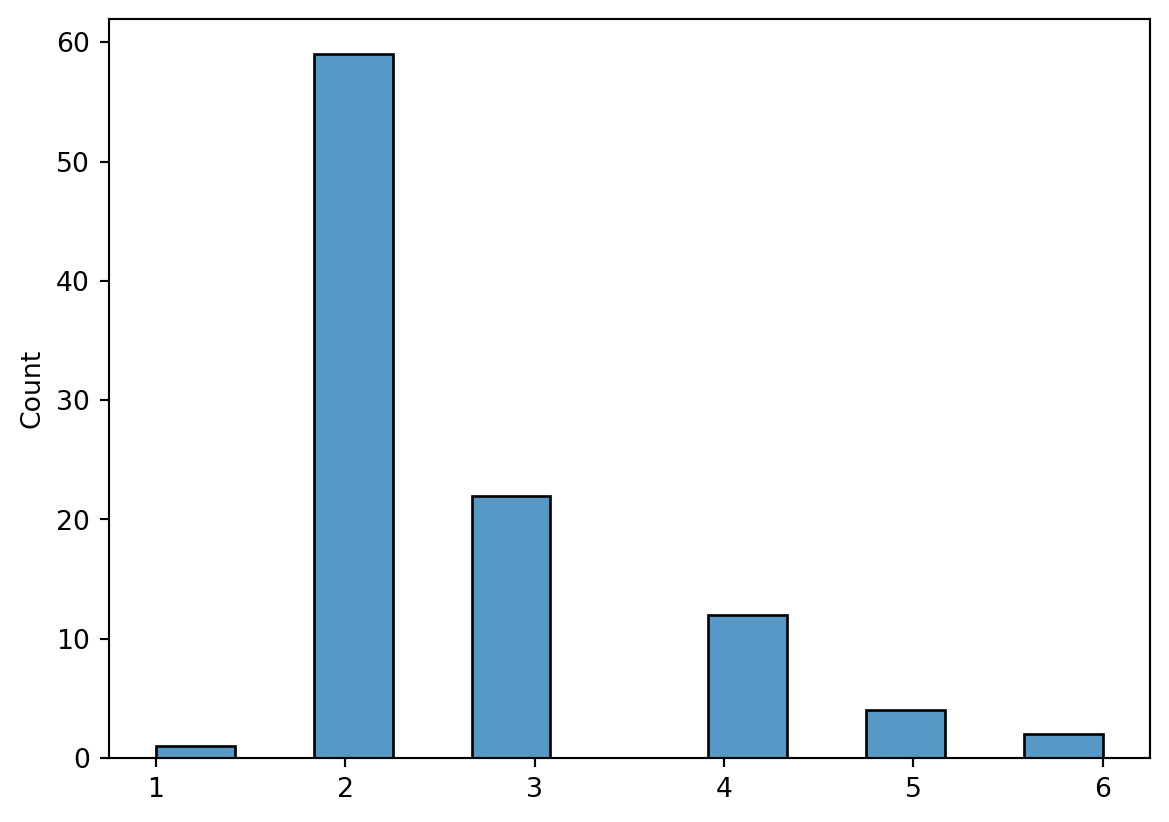
5.1 Larger
input_families = "../exp/mutual/results/20240518_TAIR10_diamond_mcl.mcl.tsv"
families_df = pd.read_csv(input_families, sep="\t", names=["gene", "family"])
families_df.head()| gene | family | |
|---|---|---|
| 0 | AT3G22150 | 1 |
| 1 | AT3G47840 | 1 |
| 2 | AT4G32430 | 1 |
| 3 | AT2G33680 | 1 |
| 4 | AT1G11290 | 1 |
families_count = families_df.groupby("family").size()
families_countfamily
1 39
2 39
3 37
4 30
5 30
..
5339 1
5340 1
5341 1
5342 1
5343 1
Length: 5343, dtype: int64sns.displot(families_count)
plt.title("Gene families size distribution in TAIR10")
plt.xlabel("Size")
plt.show()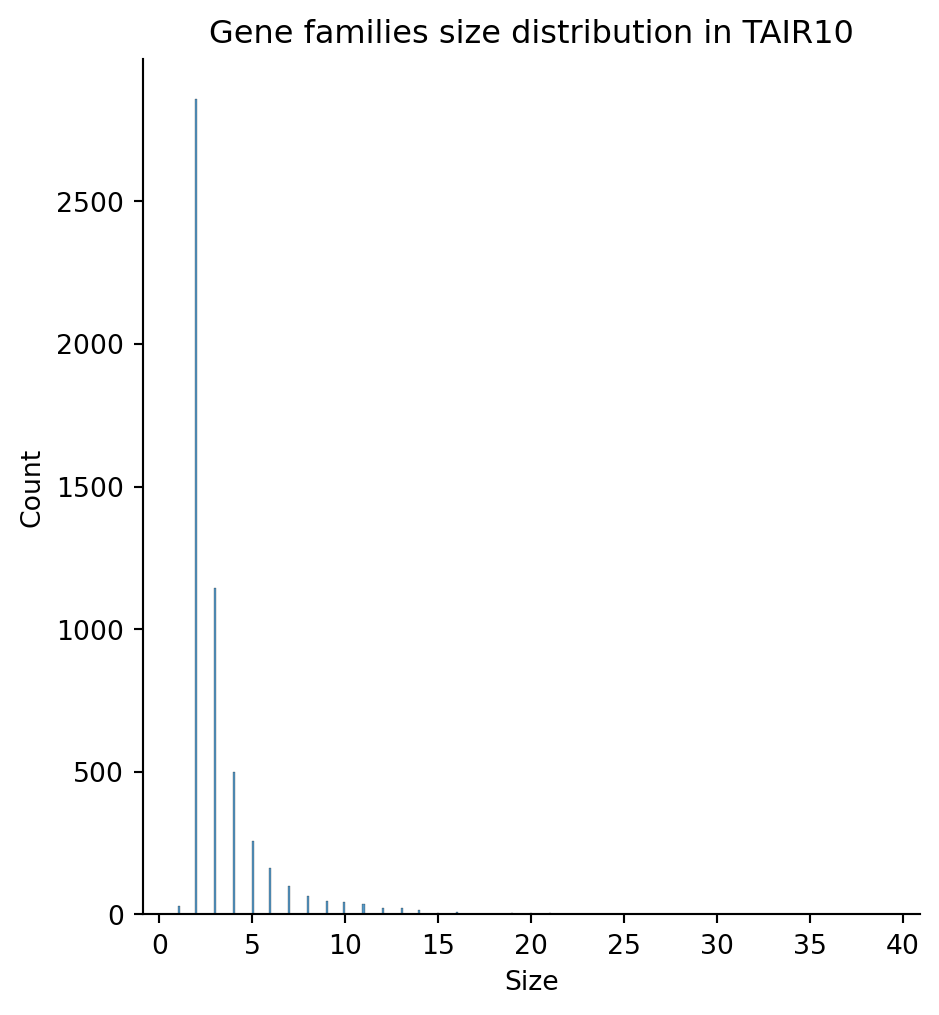
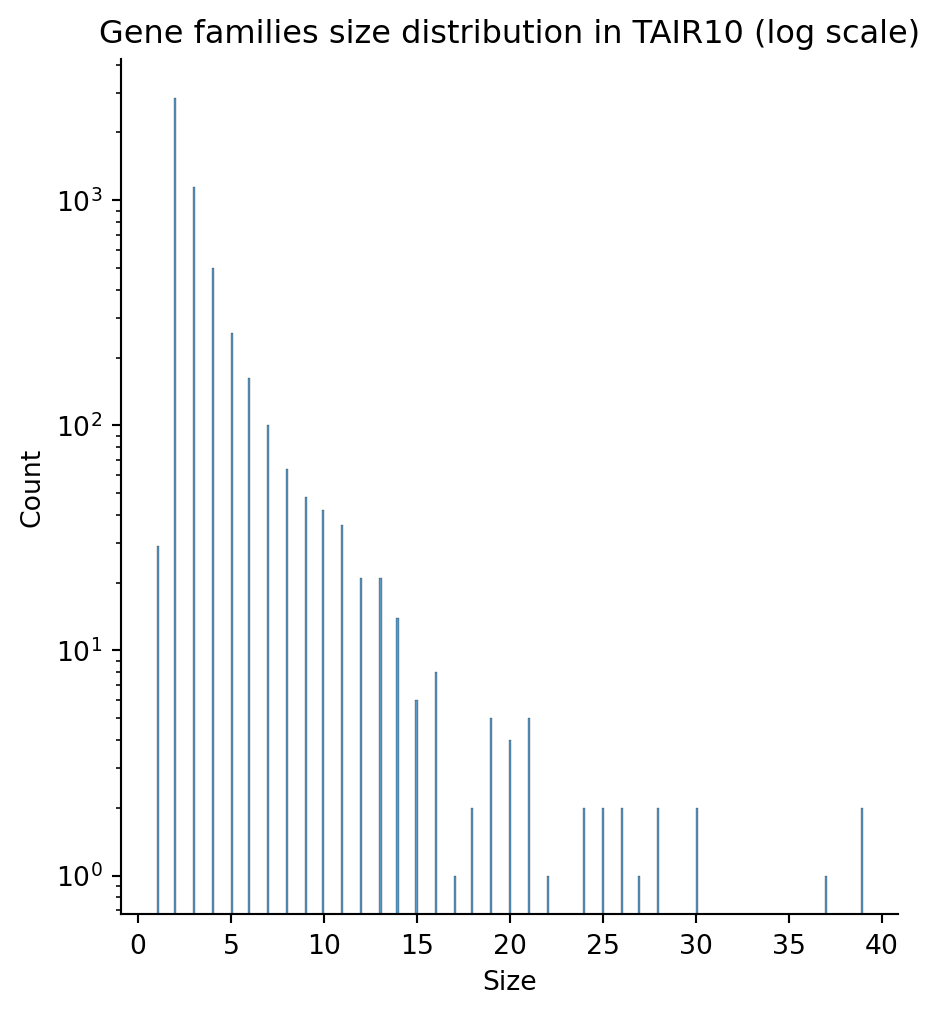
sns.displot(families_count)
plt.yscale('log')
plt.title("Gene families size distribution in TAIR10 (log scale)")
plt.xlabel("Size")
plt.show()